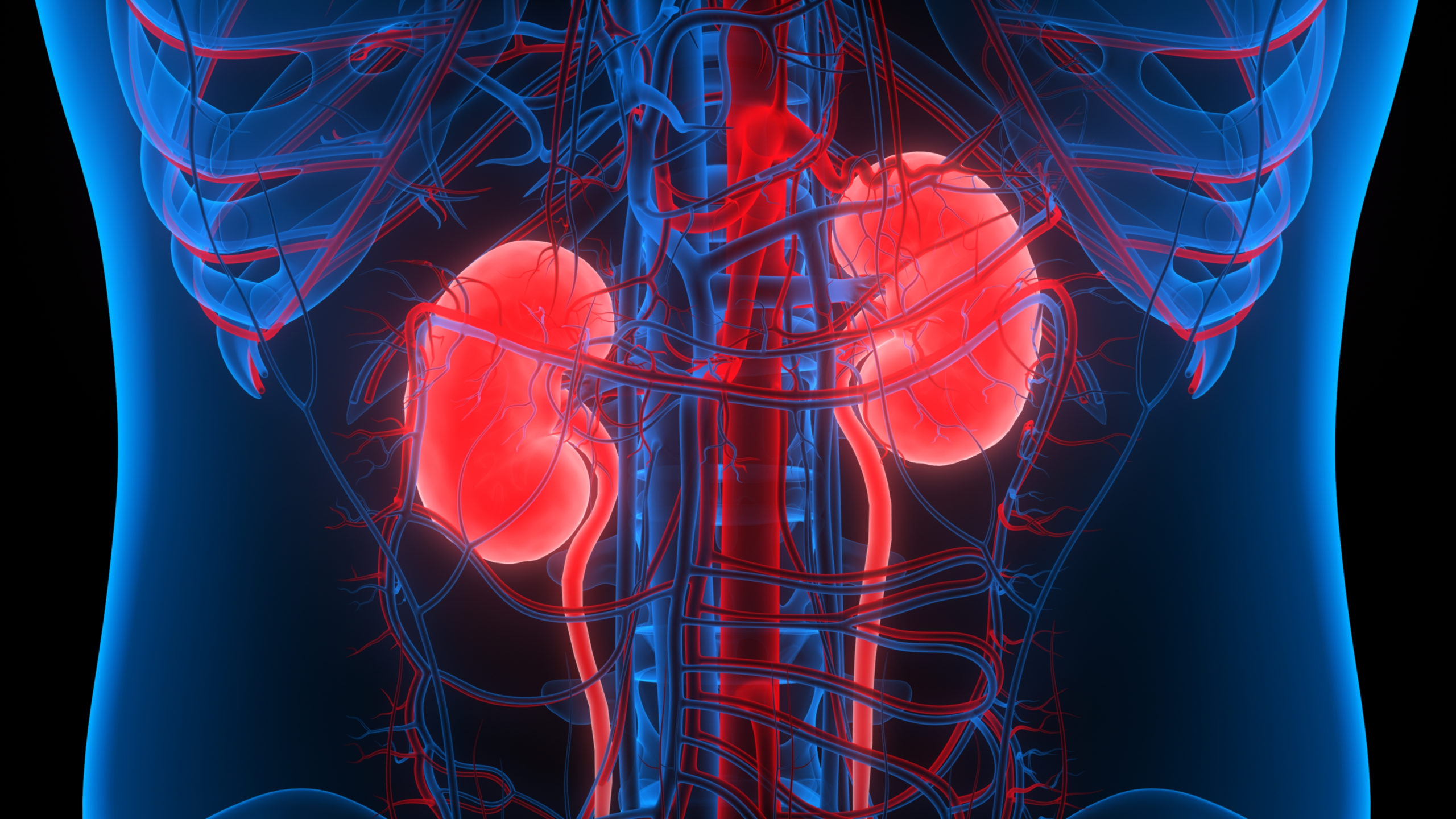
Michael Atkins, MD, of Georgetown Lombardi Comprehensive Cancer Center, and Katy Beckermann, MD, PhD, of Vanderbilt-Ingram Cancer Center, discuss the rationale behind targeting LAG-3 as a novel checkpoint inhibitor, highlighting its potential in combination therapy with anti-PD-1 in kidney cancer.
Together, they address questions regarding the durability of responses compared to CTLA-4 inhibitors and discusses ongoing research and considerations for future treatment strategies.